How does ADPS™ achieve ultra-high
detection sensitivity?
The key to the highest detection sensitivity is ADPS™ smart DNA polymerase
GENECAST has developed a new DNA polymerase that is directly involved in gene amplification, thus solving the limitations of detection sensitivity and false positives which is a common problem in ctDNA liquid biopsy cancer diagnosis. The new ADPS™ smart DNA polymerase is equipped with the intrinsic ability to distinguish between normal and mutant genes, amplifying only the mutant gene while leaving the normal gene intact.
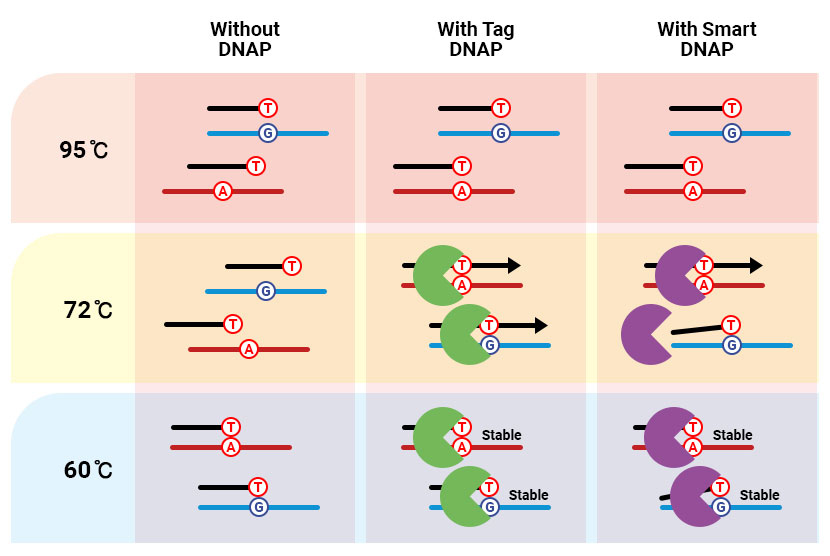
The effect of ADPS™ smart DNA polymerase on allele discrimination
Features of ADPS™ smart DNA polymerase
DNA polymerase is an enzyme that recognizes and amplifies the DNA template-primer complex. ADPS™ smart DNA polymerase can distinguish between a primer being perfectly annealed on a template and a 3'-end mismatch. This innovative discriminating ability drastically improves the detection sensitivity of mutant genes by enabling selective gene amplification. The enhanced detection sensitivity of ADPS™ smart DNA polymerase facilitates the accurate analysis of ctDNA from patients who have early cancer. Furthermore, ADPS™ smart DNA polymerase also can be used in various molecular diagnostic procedures such as NIPT and transplantation tests that require discernment.
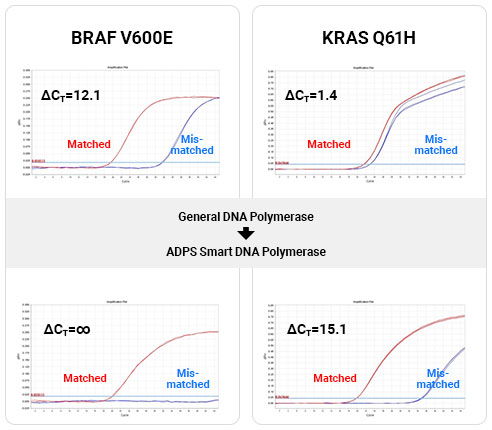
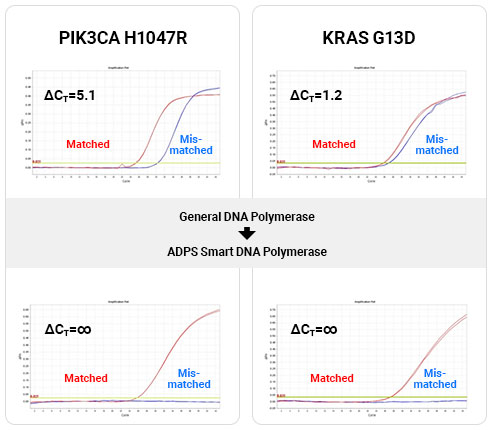
ADPS™ smart DNA polymerase versus conventional DNA polymerase qPCR experiment
Maximum detection sensitivity 0.0001%
ADPS™ smart DNA polymerase achieved a maximum detection sensitivity of 0.0001%. Through experimentation involving the BRAF V600E mutant, it was confirmed that the mutant can be quantitatively analyzed in the range of MAFs 100% to 0.0001% based on 3 x 106 copies.
| Total Number of Gene | BRAF V600E | Mutant Allele Fraction (%) | Ct |
|---|---|---|---|
| 3x106 | 3x106 | 100 | 17.5 |
| 3x106 | 3x105 | 10 | 20.8 |
| 3x106 | 3x104 | 1 | 24.5 |
| 3x106 | 3x103 | 0.1 | 28.1 |
| 3x106 | 3x102 | 0.01 | 31.8 |
| 3x106 | 3x101 | 0.001 | 35.7 |
| 3x106 | 3x100 | 0.0001 | 39.4 |
| NTC (0%) | ND |
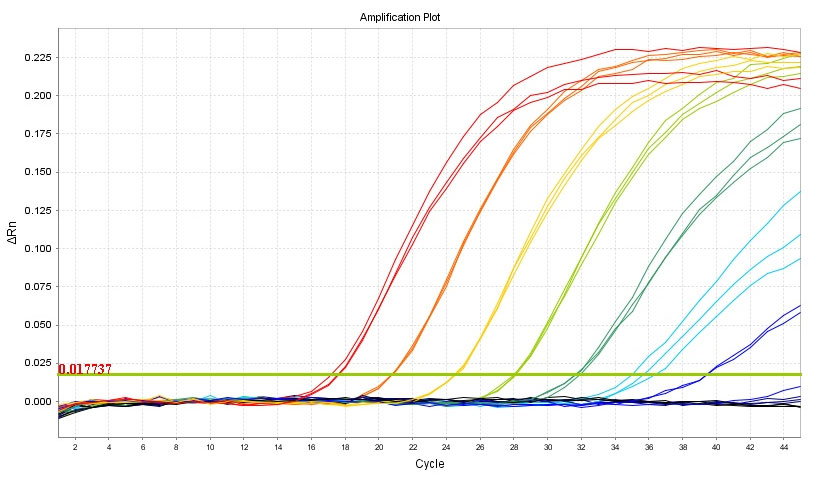
Experimental data on ADPS™ smart DNA polymerase maximum detection sensitivity
Using the ADPS™ smart DNA polymerase ADPS™ can precisely analyze cancer genes by selectively amplifying them while leaving the normal genes intact. GENECAST’s technology provides a detection sensitivity of 0.0001% without false positives.
See “How dose ADPS™ function?” “What is ADPS™?” to learn more about ADPS™.